Mastering the EM Algorithm: A Complete Guide with Mathematical Derivations and Python Implementation
The Expectation-Maximization (EM) Algorithm: A Complete Guide The Expectation-Maximization (EM) algorithm is one of the most elegant and powerful techniques in machine learning for dealing with...
# The Expectation-Maximization (EM) Algorithm: A Complete Guide
The Expectation-Maximization (EM) algorithm is one of the most elegant and powerful techniques in machine learning for dealing with incomplete data and latent variables. If you've ever wondered how Gaussian Mixture Models work, how to handle missing data gracefully, or how machines can discover hidden patterns without labels — you're in the right place.
# What is the EM Algorithm?
The Expectation-Maximization (EM) algorithm is an iterative method for finding maximum likelihood estimates of parameters in statistical models where:
- The data is incomplete or has missing values, or
- The model depends on unobserved latent variables
**Analogy:** Imagine you have a bag of coins, but you don't know which coins are fair and which are biased. You flip them and record the results, but someone forgot to label which coin produced which result. The EM algorithm helps you figure out **both** which coin is which **AND** what their bias probabilities are — simultaneously!
# The Core Idea
The brilliance of EM lies in its iterative two-step process:
- E-Step (Expectation): Given your current parameter estimates, calculate the expected value of the log-likelihood function with respect to the conditional distribution of the latent variables.
- M-Step (Maximization): Find the parameters that maximize the expected log-likelihood computed in the E-step.
You repeat these steps until convergence — when the parameters stop changing significantly.
# Mathematical Foundation
# The Setup
Let's formalize this. Suppose we have:
| Symbol | Description |
|---|---|
| X | Observed data (what we can see) |
| Z | Hidden/latent variables (what we can't see) |
| \theta | Parameters we want to estimate |
| P(X, Z \mid $\theta$) | Joint probability of observed and hidden data |
Our goal is to maximize the log-likelihood of the observed data:
But computing P(X \theta$) requires marginalizing over $Z:
This summation (or integral for continuous $Z$) is often intractable. Enter EM!
# Jensen's Inequality and the Lower Bound
Starting with the log-likelihood:
We introduce an arbitrary distribution over the latent variables:
By Jensen's inequality (since log is concave):
This right-hand side is called the Evidence Lower Bound (ELBO):
Which expands to:
where is the entropy of .
# E-Step: Expectation
Fix to its current estimate and choose to maximize the lower bound. The optimal choice is:
This is the posterior distribution of the latent variables given the observed data and current parameters.
# M-Step: Maximization
Fix and maximize the lower bound with respect to :
This is typically easier than the original problem because the log can now be pushed inside the expectation.
# Convergence Guarantee
Each iteration of EM is guaranteed to increase the log-likelihood (or keep it the same):
This monotonic increase ensures convergence to at least a local maximum.
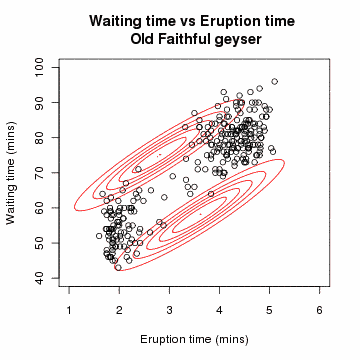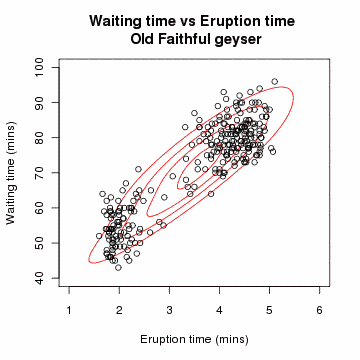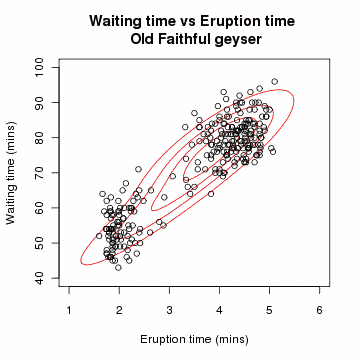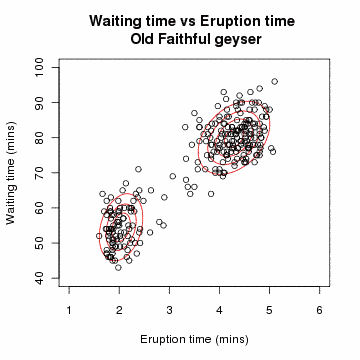
# Gaussian Mixture Model: A Concrete Example
# Problem Setup
Suppose we have data points from different Gaussian distributions, but we don't know which point came from which Gaussian.
Data: where
Latent variable: — which component generated
Parameters:
- : Mixing coefficients, where
- : Mean of component
- : Covariance matrix of component
Generative Process:
- Choose component with probability
- Sample
# E-Step: Computing Responsibilities
where the Gaussian density is:
These values are called responsibilities.
# M-Step: Updating Parameters
Mixing coefficients:
Means:
Covariances:
# Numerical Example: GMM with 1D Data
# Setup
Data: , suspected from Gaussians.
Initialization:
| Parameter | Value |
|---|---|
| \pi_1^{(0)}, \pi_2^{(0)} | 0.5,\ 0.5 |
| \mu_1^{(0)}, \mu_2^{(0)} | 2.0,\ 7.0 |
| \sigma_1^{(0)}, \sigma_2^{(0)} | 1.0,\ 1.0 |
# Iteration 1 — E-Step
For :
Full responsibility table:
| Point | x_i | \gamma_{i1} | \gamma_{i2} |
|---|---|---|---|
| 1 | 1.0 | 0.99998 | 0.00002 |
| 2 | 1.5 | 0.9953 | 0.0047 |
| 3 | 2.0 | 0.9502 | 0.0498 |
| 4 | 8.0 | 0.0018 | 0.9982 |
| 5 | 8.5 | 0.0001 | 0.9999 |
| 6 | 9.0 | 0.00001 | 0.99999 |
# Iteration 1 — M-Step
After convergence:
| Cluster | \mu | \sigma | \pi |
|---|---|---|---|
| 1 | ≈ 1.5 | ≈ 0.5 | ≈ 0.5 |
| 2 | ≈ 8.5 | ≈ 0.5 | ≈ 0.5 |
# Python Implementation
# 1D GMM from Scratch
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import norm
class GaussianMixtureEM:
"""Gaussian Mixture Model using the EM Algorithm (1D)."""
def __init__(self, n_components=2, max_iter=100, tol=1e-4):
self.n_components = n_components
self.max_iter = max_iter
self.tol = tol
def _initialize_parameters(self, X):
n_samples = X.shape
random_idx = np.random.choice(n_samples, self.n_components, replace=False)
self.means_ = X[random_idx]
self.covariances_ = np.array([np.var(X)] * self.n_components)
self.weights_ = np.ones(self.n_components) / self.n_components
def _e_step(self, X):
responsibilities = np.zeros((X.shape, self.n_components))
for k in range(self.n_components):
responsibilities[:, k] = self.weights_[k] * norm.pdf(
X, self.means_[k], np.sqrt(self.covariances_[k])
)
responsibilities /= responsibilities.sum(axis=1, keepdims=True)
return responsibilities
def _m_step(self, X, responsibilities):
Nk = responsibilities.sum(axis=0)
self.means_ = (responsibilities.T @ X) / Nk
for k in range(self.n_components):
diff = X - self.means_[k]
self.covariances_[k] = (responsibilities[:, k] * diff**2).sum() / Nk[k]
self.weights_ = Nk / X.shape
def _compute_log_likelihood(self, X):
log_likelihood = 0
for i in range(X.shape):
sample_ll = sum(
self.weights_[k] * norm.pdf(X[i], self.means_[k], np.sqrt(self.covariances_[k]))
for k in range(self.n_components)
)
log_likelihood += np.log(sample_ll)
return log_likelihood
def fit(self, X):
X = np.array(X).reshape(-1)
self._initialize_parameters(X)
log_likelihood_old = -np.inf
self.log_likelihoods_ = []
for iteration in range(self.max_iter):
responsibilities = self._e_step(X)
self._m_step(X, responsibilities)
log_likelihood = self._compute_log_likelihood(X)
self.log_likelihoods_.append(log_likelihood)
if abs(log_likelihood - log_likelihood_old) < self.tol:
print(f"Converged at iteration {iteration + 1}")
break
log_likelihood_old = log_likelihood
return self
def predict_proba(self, X):
return self._e_step(np.array(X).reshape(-1))
def predict(self, X):
return self.predict_proba(X).argmax(axis=1)
# --- Example Usage ---
if __name__ == "__main__":
np.random.seed(42)
data1 = np.random.normal(1.5, 0.5, 300)
data2 = np.random.normal(8.5, 0.5, 300)
X = np.concatenate([data1, data2])
np.random.shuffle(X)
gmm = GaussianMixtureEM(n_components=2, max_iter=100)
gmm.fit(X)
print(f"Means: {gmm.means_}")
print(f"Std Devs: {np.sqrt(gmm.covariances_)}")
print(f"Weights: {gmm.weights_}")
labels = gmm.predict(X)
fig, axes = plt.subplots(2, 2, figsize=(14, 10))
x_range = np.linspace(X.min(), X.max(), 1000)
# Plot 1: Histogram + fitted components
ax = axes
ax.hist(X, bins=50, density=True, alpha=0.6, color="gray", label="Data")
for k in range(gmm.n_components):
component = gmm.weights_[k] * norm.pdf(x_range, gmm.means_[k], np.sqrt(gmm.covariances_[k]))
ax.plot(x_range, component, label=f"Component {k+1}", linewidth=2)
mixture = sum(
gmm.weights_[k] * norm.pdf(x_range, gmm.means_[k], np.sqrt(gmm.covariances_[k]))
for k in range(gmm.n_components)
)
ax.plot(x_range, mixture, "r--", linewidth=2, label="Mixture")
ax.set(xlabel="Value", ylabel="Density", title="GMM Fit to Data")
ax.legend(); ax.grid(alpha=0.3)
# Plot 2: Cluster assignments
ax = axes[1]
for k in range(gmm.n_components):
cluster_data = X[labels == k]
ax.scatter(cluster_data, np.zeros_like(cluster_data) + k, alpha=0.6, s=30, label=f"Cluster {k+1}")
ax.set(xlabel="Value", ylabel="Cluster", title="Cluster Assignments")
ax.set_yticks(range(gmm.n_components))
ax.legend(); ax.grid(alpha=0.3)
# Plot 3: Log-likelihood convergence
ax = axes[1]
ax.plot(gmm.log_likelihoods_, linewidth=2, color="blue")
ax.set(xlabel="Iteration", ylabel="Log-Likelihood", title="EM Algorithm Convergence")
ax.grid(alpha=0.3)
# Plot 4: Responsibilities for sampled points
ax = axes[1]
sample_indices = np.linspace(0, len(X) - 1, 10, dtype=int)
responsibilities = gmm.predict_proba(X[sample_indices])
x_pos = np.arange(len(sample_indices))
width = 0.35
for k in range(gmm.n_components):
ax.bar(x_pos + k * width, responsibilities[:, k], width, label=f"Component {k+1}", alpha=0.8)
ax.set(xlabel="Sample Index", ylabel="Responsibility", title="Posterior Probabilities (Responsibilities)")
ax.set_xticks(x_pos + width / 2)
ax.set_xticklabels(sample_indices)
ax.legend(); ax.grid(alpha=0.3, axis="y")
plt.tight_layout()
plt.savefig("em_algorithm_results.png", dpi=300, bbox_inches="tight")
plt.show()# 2D GMM
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.patches import Ellipse
from scipy.stats import multivariate_normal
class GaussianMixture2D:
"""2D Gaussian Mixture Model with EM."""
def __init__(self, n_components=3, max_iter=100, tol=1e-4):
self.n_components = n_components
self.max_iter = max_iter
self.tol = tol
def _initialize_parameters(self, X):
n_samples, n_features = X.shape
random_idx = np.random.choice(n_samples, self.n_components, replace=False)
self.means_ = X[random_idx]
self.covariances_ = np.array([np.eye(n_features)] * self.n_components)
self.weights_ = np.ones(self.n_components) / self.n_components
def _e_step(self, X):
responsibilities = np.zeros((X.shape, self.n_components))
for k in range(self.n_components):
responsibilities[:, k] = self.weights_[k] * multivariate_normal.pdf(
X, self.means_[k], self.covariances_[k]
)
responsibilities /= responsibilities.sum(axis=1, keepdims=True)
return responsibilities
def _m_step(self, X, responsibilities):
n_samples, n_features = X.shape
Nk = responsibilities.sum(axis=0)
self.means_ = (responsibilities.T @ X) / Nk[:, np.newaxis]
for k in range(self.n_components):
diff = X - self.means_[k]
self.covariances_[k] = (responsibilities[:, k, np.newaxis] * diff).T @ diff / Nk[k]
self.covariances_[k] += 1e-6 * np.eye(n_features) # regularization
self.weights_ = Nk / n_samples
def _compute_log_likelihood(self, X):
log_likelihood = 0
for i in range(X.shape):
sample_ll = sum(
self.weights_[k] * multivariate_normal.pdf(X[i], self.means_[k], self.covariances_[k])
for k in range(self.n_components)
)
log_likelihood += np.log(sample_ll + 1e-10)
return log_likelihood
def fit(self, X):
X = np.array(X)
self._initialize_parameters(X)
log_likelihood_old = -np.inf
self.log_likelihoods_ = []
for iteration in range(self.max_iter):
responsibilities = self._e_step(X)
self._m_step(X, responsibilities)
log_likelihood = self._compute_log_likelihood(X)
self.log_likelihoods_.append(log_likelihood)
if abs(log_likelihood - log_likelihood_old) < self.tol:
print(f"Converged at iteration {iteration + 1}")
break
log_likelihood_old = log_likelihood
return self
def predict(self, X):
return self._e_step(X).argmax(axis=1)
# --- Example Usage ---
if __name__ == "__main__":
np.random.seed(42)
cluster1 = np.random.multivariate_normal(, [[1, 0.5], [0.5, 1]], 200)
cluster2 = np.random.multivariate_normal(, [[1, -0.3], [-0.3, 1]], 200)[3]
cluster3 = np.random.multivariate_normal(, [[1.5, 0], [0, 0.5]], 200)[4][1]
X_2d = np.vstack([cluster1, cluster2, cluster3])
np.random.shuffle(X_2d)
gmm_2d = GaussianMixture2D(n_components=3, max_iter=100)
gmm_2d.fit(X_2d)
labels_2d = gmm_2d.predict(X_2d)
fig, axes = plt.subplots(1, 2, figsize=(16, 6))
ax = axes
ax.scatter(X_2d[:, 0], X_2d[:, 1], c=labels_2d, cmap="viridis",
alpha=0.6, s=30, edgecolors="k", linewidth=0.5)
ax.scatter(gmm_2d.means_[:, 0], gmm_2d.means_[:, 1],
c="red", marker="X", s=300, edgecolors="black", linewidth=2, label="Cluster Centers")
for k in range(gmm_2d.n_components):
eigenvalues, eigenvectors = np.linalg.eigh(gmm_2d.covariances_[k])
angle = np.degrees(np.arctan2(eigenvectors, eigenvectors))[1]
for scale in:[5][1]
ellipse = Ellipse(
gmm_2d.means_[k],
width=2 * scale * np.sqrt(eigenvalues),
height=2 * scale * np.sqrt(eigenvalues),[1]
angle=angle, facecolor="none", edgecolor="red",
linewidth=2, linestyle="--", alpha=0.7,
)
ax.add_patch(ellipse)
ax.set(xlabel="$X_1$", ylabel="$X_2$", title="2D GMM Clustering Results")
ax.legend(); ax.grid(alpha=0.3)
ax = axes[1]
ax.plot(gmm_2d.log_likelihoods_, linewidth=3, color="darkblue")
ax.set(xlabel="Iteration", ylabel="Log-Likelihood", title="EM Convergence for 2D GMM")
ax.grid(alpha=0.3)
plt.tight_layout()
plt.savefig("em_2d_gmm_results.png", dpi=300, bbox_inches="tight")
plt.show()
print(f"Means:\n{gmm_2d.means_}")
print(f"Weights: {gmm_2d.weights_}")# Key Code Components
| Component | Description |
|---|---|
| _initialize_parameters | Randomly picks data points as means; sets uniform weights and data variance for covariances |
| _e_step | Computes responsibilities via Bayes' theorem |
| _m_step | Updates \mu_k, \Sigma_k, \pi_k using weighted MLE |
| _compute_log_likelihood | Tracks monotonic increase for convergence monitoring |
| predict / predict_proba | Assigns hard or soft cluster labels |
# Applications
# Clustering & Density Estimation
- Gaussian Mixture Models — soft clustering with probabilistic assignments
- Hidden Markov Models — speech recognition, NLP
- Topic Modeling — Latent Dirichlet Allocation (LDA)
# Missing Data Imputation
- Filling in missing values in datasets
- Recommendation systems with incomplete user-item matrices
- Medical records with missing patient information
# Computer Vision
- Image segmentation and object tracking
- Background subtraction in video
# Bioinformatics & Finance
- Gene expression analysis, protein structure prediction
- Market regime detection, credit risk modeling
# Advantages and Limitations
| Aspect | Details |
|---|---|
| ✅ Principled | Based on maximum likelihood with solid theoretical guarantees |
| ✅ Handles latent variables | Naturally incorporates unobserved structure |
| ✅ Guaranteed convergence | Monotonically increases log-likelihood each iteration |
| ✅ Soft assignments | Returns probabilities, not just hard cluster labels |
| ❌ Local maxima | Only guaranteed to find a local, not global, maximum |
| ❌ Initialization sensitive | Poor starts lead to poor solutions |
| ❌ K must be specified | Number of components chosen by the user |
| ❌ Slow on large data | Quadratic scaling with dimension in full-covariance models |
# Tips for Effective Use
# Initialization Strategies
# Option 1: Random initialization
random_idx = np.random.choice(n_samples, n_components)
means = X[random_idx]
# Option 2: K-Means++ warm start (recommended)
from sklearn.cluster import KMeans
kmeans = KMeans(n_clusters=n_components, init="k-means++")
kmeans.fit(X)
means = kmeans.cluster_centers_
# Option 3: Multiple restarts
best_gmm, best_ll = None, -np.inf
for _ in range(n_restarts):
gmm = GaussianMixtureEM(n_components=k)
gmm.fit(X)
if gmm.log_likelihoods_[-1] > best_ll:
best_gmm = gmm
best_ll = gmm.log_likelihoods_[-1]# Model Selection with AIC / BIC
Use information criteria to select the number of components :
where is the likelihood, is the number of parameters, and is the number of samples.
def compute_bic(gmm, X):
n_params = gmm.n_components * 3 - 1 # means, variances, weights
return -2 * gmm.log_likelihoods_[-1] + n_params * np.log(len(X))
bic_scores = []
for k in range(1, 10):
gmm = GaussianMixtureEM(n_components=k)
gmm.fit(X)
bic_scores.append(compute_bic(gmm, X))
optimal_k = np.argmin(bic_scores) + 1# Regularization for Numerical Stability
# Add small diagonal term to prevent singular covariance matrices
self.covariances_[k] += 1e-6 * np.eye(n_features)# Convergence Monitoring
converged = (
abs(log_likelihood - log_likelihood_old) < tol
or np.allclose(means_old, self.means_, atol=tol)
)# Algorithm Comparison
| Algorithm | Type | Advantages | Disadvantages |
|---|---|---|---|
| EM / GMM | Soft clustering | Probabilistic, handles uncertainty | Local maxima, K must be specified |
| K-Means | Hard clustering | Fast, simple | Hard assignments, assumes spherical clusters |
| DBSCAN | Density-based | Finds arbitrary shapes, auto-detects K | Sensitive to \varepsilon and min_samples |
| Hierarchical | Agglomerative | No K needed, dendrogram visualization | O(n^2) memory, no probability output |
# Further Reading
- Dempster, A. P., Laird, N. M., & Rubin, D. B. (1977). Maximum likelihood from incomplete data via the EM algorithm. Journal of the Royal Statistical Society, Series B, 39(1), 1–38.
- Bishop, C. M. (2006). Pattern Recognition and Machine Learning. Springer. Chapter 9.
- Murphy, K. P. (2012). Machine Learning: A Probabilistic Perspective. MIT Press. Chapter 11.

About the Author
Dr. Himanshu Rai is an Assistant Professor in Computer Science & Engineering at SRM Institute of Science and Technology, specializing in Machine Learning, Artificial Intelligence, and Data Science. She is passionate about advancing research in computational intelligence and mentoring students.
